|
Mathematics
Is Biology's Next Microscope, Only Better; Biology Is Mathematics' Next
Physics, Only Better Joel
E. Cohen is at the Laboratory of Populations, Rockefeller and Columbia Universities,
New York, New York, United States of America. E-mail: cohen@rockefeller.edu Copyright: © 2004 Joel E. Cohen. This is an open-access article
distributed under the terms of the Creative Commons Attribution License,
which permits unrestricted use, distribution, and reproduction in any medium,
provided the original work is properly cited. This work appeared in PLoS Biol
2(12): e439. http://www.plosbiology.org/ Although
mathematics has long been intertwined with the biological sciences, an
explosive synergy between biology and mathematics seems poised to enrich and
extend both fields greatly in the coming decades (Levin 1992; Murray 1993;
Jungck 1997; Hastings et al. 2003; Palmer
et al. 2003; Hastings and Palmer 2003).
Biology will increasingly stimulate the creation of qualitatively new realms
of mathematics. Why? In biology, ensemble properties emerge at each level of
organization from the interactions of heterogeneous biological units at that
level and at lower and higher levels of organization (larger and smaller
physical scales, faster and slower temporal scales). New mathematics will be
required to cope with these ensemble properties and with the heterogeneity of
the biological units that compose ensembles at each level. The discovery of
the microscope in the late 17th century caused a revolution in biology by
revealing otherwise invisible and previously unsuspected worlds. Western
cosmology from classical times through the end of the Renaissance envisioned
a system with three types of spheres: the sphere of man, exemplified by his
imperfectly round head; the sphere of the world, exemplified by the
imperfectly spherical earth; and the eight perfect spheres of the universe,
in which the seven (then known) planets moved and the outer stars were fixed
(Nicolson 1960). The discovery of a
microbial world too small to be seen by the naked eye challenged the
completeness of this cosmology and unequivocally demonstrated the existence
of living creatures unknown to the Scriptures of Old World religions. Mathematics
broadly interpreted is a more general microscope. It can reveal otherwise
invisible worlds in all kinds of data, not only optical. For example,
computed tomography can reveal a cross-section of a human head from the
density of X-ray beams without ever opening the head, by using the Radon
transform to infer the densities of materials at each location within the head
(Hsieh 2003). Charles Darwin was right
when he wrote that people with an understanding “of the great leading
principles of mathematics… seem to have an extra sense” (F. Darwin 1905). Today's biologists increasingly recognize
that appropriate mathematics can help interpret any kind of data. In this
sense, mathematics is biology's next microscope, only better. Conversely,
mathematics will benefit increasingly from its involvement with biology, just
as mathematics has already benefited and will continue to benefit from its
historic involvement with physical problems. In classical times, physics, as
first an applied then a basic science, stimulated enormous advances in
mathematics. For example, geometry reveals by its very etymology (geometry)
its origin in the needs to survey the lands and waters of Earth. Geometry was
used to lay out fields in Egypt after the flooding of the Nile, to aid
navigation, to aid city planning. The inventions of the calculus by Isaac
Newton and Gottfried Leibniz in the later 17th century were stimulated by
physical problems such as planetary orbits and optical calculations. In the coming
century, biology will stimulate the creation of entirely new realms of
mathematics. In this sense, biology is mathematics' next physics, only
better. Biology will stimulate fundamentally new mathematics because living
nature is qualitatively more heterogeneous than non-living nature. For
example, it is estimated that there are 2,000–5,000 species of rocks and
minerals in the earth's crust, generated from the hundred or so naturally
occurring elements (Shipman et al. 2003;
chapter 21 estimates 2,000 minerals in Earth's crust). By contrast, there are
probably between 3 million and 100 million biological species on Earth,
generated from a small fraction of the naturally occurring elements. If
species of rocks and minerals may validly be compared with species of living
organisms, the living world has at least a thousand times the diversity of
the non-living. This comparison omits the enormous evolutionary importance of
individual variability within species. Coping with the hyper-diversity of
life at every scale of spatial and temporal organization will require
fundamental conceptual advances in mathematics. The Past The interactions
between mathematics and biology at present follow from their interactions
over the last half millennium. The discovery of the New World by Europeans
approximately 500 years ago—and of its many biological species not described
in religious Scriptures—gave impetus to major conceptual progress in biology. The outstanding
milestone in the early history of biological quantification was the work of William
Harvey, Exercitatio Anatomica De Motu Cordis et Sanguinis In Animalibus
(An Anatomical Disquisition on the Motion of the Heart and Blood in Animals)
(Harvey 1847), first published in 1628. Harvey's demonstration that the blood
circulates was the pivotal founding event of the modern interaction between
mathematics and biology. His elegant reasoning is worth understanding. From the time of
the ancient Greek physician Galen (131–201 C.E.) until William Harvey studied
medicine in Padua (1600–1602, while Galileo was active there), it was
believed that there were two kinds of blood, arterial blood and venous blood.
Both kinds of blood were believed to ebb and flow under the motive power of
the liver, just as the tides of the earth ebbed and flowed under the motive
power of the moon. Harvey became physician to the king of England. He used
his position of privilege to dissect deer from the king's deer park as well
as executed criminals. Harvey observed that the veins in the human arm have
one-way valves that permit blood to flow from the periphery toward the heart
but not in the reverse direction. Hence the theory that the blood ebbs and
flows in both veins and arteries could not be correct. Harvey also
observed that the heart was a contractile muscle with one-way valves between
the chambers on each side. He measured the volume of the left ventricle of
dead human hearts and found that it held about two ounces (about 60 ml),
varying from 1.5 to three ounces in different individuals. He estimated that
at least one-eighth and perhaps as much as one-quarter of the blood in the
left ventricle was expelled with each stroke of the heart. He measured that
the heart beat 60–100 times per minute. Therefore, the volume of blood
expelled from the left ventricle per hour was about 60 ml × 1/8 × 60
beats/minute × 60 minutes/hour, or 27 litres/hour. However, the average human
has only 5.5 litres of blood (a quantity that could be estimated by draining
a cadaver). Therefore, the blood must be like a stage army that marches off
one side of the stage, returns behind the scenes, and re-enters from the
other side of the stage, again and again. The large volume of blood pumped
per hour could not possibly be accounted for by the then-prevalent theory
that the blood originated from the consumption of food. Harvey inferred that
there must be some small vessels that conveyed the blood from the outgoing
arteries to the returning veins, but he was not able to see those small
vessels. His theoretical prediction, based on his meticulous anatomical
observations and his mathematical calculations, was spectacularly confirmed
more than half a century later when Marcello Malpighi (1628–1694) saw the
capillaries under a microscope. Harvey's discovery illustrates the enormous
power of simple, off-the-shelf mathematics combined with careful observation
and clear reasoning. It set a high standard for all later uses of mathematics
in biology. Mathematics was
crucial in the discovery of genes by Mendel (Orel
1984) and in the theory of evolution. Mathematics was and continues to be
the principal means of integrating evolution and genetics since the classic
work of R. A. Fisher, J. B. S. Haldane, and S. Wright in the first half of
the 20th century (Provine 2001). Over the last 500
years, mathematics has made amazing progress in each of its three major
fields: geometry and topology, algebra, and analysis. This progress has
enriched all the biological sciences. In 1637, René
Descartes linked the featureless plane of Greek geometry to the symbols and
formulas of Arabic algebra by imposing a coordinate system (conventionally, a
horizontal x-axis and a vertical y-axis) on the geometric plane and using
numbers to measure distances between points. If every biologist who plotted
data on x–y coordinates acknowledged the contribution of Descartes to
biological understanding, the key role of mathematics in biology would be
uncontested. Another highlight
of the last five centuries of geometry was the invention of non-Euclidean geometries
(1823–1830). Shocking at first, these geometries unshackled the possibilities
of mathematical reasoning from the intuitive perception of space. These
non-Euclidean geometries have made significant contributions to biology in
facilitating, for example, mapping the brain onto a flat surface (Hurdal et al. 1999; Bowers and Hurdal 2003). In algebra,
efforts to find the roots of equations led to the discovery of the symmetries
of roots of equations and thence to the invention of group theory, which
finds routine application in the study of crystallographic groups by
structural biologists today. Generalizations of single linear equations to
families of simultaneous multi-variable linear equations stimulated the
development of linear algebra and the European re-invention and naming of
matrices in the mid-19th century. The use of a matrix of numbers to solve
simultaneous systems of linear equations can be traced back in Chinese
mathematics to the period from 300 B.C.E. to 200 C.E. (in a work by Chiu
Chang Suan Shu called Nine Chapters of the Mathematical Art; Smoller 2001). In the 19th century,
matrices were considered the epitome of useless mathematical abstraction.
Then, in the 20th century, it was discovered, for example, that the numerical
processes required for the cohort-component method of population projection
can be conveniently summarized and executed using matrices (Keyfitz 1968). Today the use of matrices is routine in
agencies responsible for making official population projections as well as in
population-biological research on human and nonhuman populations (Caswell 2001). Finally, analysis,
including the calculus of Newton and Leibniz and probability theory, is the
line between ancient thought and modern thought. Without an understanding of
the concepts of analysis, especially the concept of a limit, it is not
possible to grasp much of modern science, technology, or economic theory.
Those who understand the calculus, ordinary and partial differential
equations, and probability theory have a way of seeing and understanding the
world, including the biological world, that is unavailable to those who do
not. Conceptual and
scientific challenges from biology have enriched mathematics by leading to
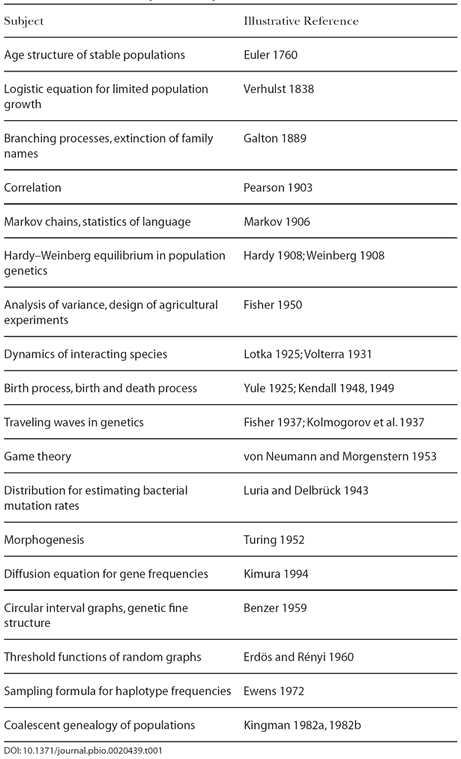
innovative thought about new kinds of mathematics. Table 1 lists examples of
new and useful mathematics arising from problems in the life sciences broadly
construed, including biology and some social sciences. Many of these
developments blend smoothly into their antecedents and later elaborations.
For example, game theory has a history before the work of John von Neumann (von Neumann 1959; von Neumann and Morgenstern 1953), and
Karl Pearson's development of the correlation coefficient (Pearson and Lee 1903) rested on earlier work by Francis Galton (1889).
The Present To see how the
interactions of biology and mathematics may proceed in the future, it is
helpful to map the present landscapes of biology and applied mathematics. The biological
landscape may be mapped as a rectangular table with different rows for
different questions and different columns for different biological domains.
Biology asks six kinds of questions. How is it built? How does it work? What
goes wrong? How is it fixed? How did it begin? What is it for? These are
questions, respectively, about structures, mechanisms, pathologies, repairs,
origins, and functions or purposes. The former teleological interpretation of
purpose has been replaced by an evolutionary perspective. Biological domains,
or levels of organization, include molecules, cells, tissues, organs,
individuals, populations, communities, ecosystems or landscapes, and the
biosphere. Many biological research problems can be classified as the
combination of one or more questions directed to one or more domains. In addition,
biological research questions have important dimensions of time and space.
Timescales of importance to biology range from the extremely fast processes
of photosynthesis to the billions of years of living evolution on Earth.
Relevant spatial scales range from the molecular to the cosmic (cosmic rays
may have played a role in evolution on Earth). The questions and the domains
of biology behave differently on different temporal and spatial scales. The opportunities
and the challenges that biology offers mathematics arise because the units at
any given level of biological organization are heterogeneous, and the
outcomes of their interactions (sometimes called “emergent phenomena” or
“ensemble properties”) on any selected temporal and spatial scale may be
substantially affected by the heterogeneity and interactions of biological
components at lower and higher levels of biological organization and at
smaller and larger temporal and spatial scales (Anderson 1972, 1995). The landscape of
applied mathematics is better visualized as a tetrahedron (a pyramid with a
triangular base) than as a matrix with temporal and spatial dimensions.
(Mathematical imagery, such as a tetrahedron for applied mathematics and a
matrix for biology, is useful even in trying to visualize the landscapes of
biology and mathematics.) The four main points of the applied mathematical
landscape are data structures, algorithms, theories and models (including all
pure mathematics), and computers and software. Data structures are ways to
organize data, such as the matrix used above to describe the biological
landscape. Algorithms are procedures for manipulating symbols. Some
algorithms are used to analyse data, others to analyse models. Theories and
models, including the theories of pure mathematics, are used to analyse both
data and ideas. Mathematics and mathematical theories provide a testing
ground for ideas in which the strength of competing theories can be measured.
Computers and software are an important, and frequently the most visible,
vertex of the applied mathematical landscape. However, cheap, easy computing
increases the importance of theoretical understanding of the results of
computation. Theoretical understanding is required as a check on the great
risk of error in software, and to bridge the enormous gap between
computational results and insight or understanding. The landscape of
research in mathematics and biology contains all combinations of one or more
biological questions, domains, time scales, and spatial scales with one or
more data structures, algorithms, theories or models, and means of
computation (typically software and hardware). The following example from
cancer biology illustrates such a combination: the question, “how does it
work?” is approached in the domain of cells (specifically, human cancer
cells) with algorithms for correlation and hierarchical clustering. Gene
expression and drug activity in human cancer. Suppose a person
has a cancer. Could information about the activities of the genes in the
cells of the person's cancer guide the use of cancer-treatment drugs so that
more effective drugs are used and less effective drugs are avoided? To
suggest answers to this question, Scherf et
al. (2000) ingeniously applied off-the-shelf mathematics, specifically,
correlation—invented nearly a century earlier by Karl Pearson (Pearson and Lee 1903) in a study of human
inheritance—and clustering algorithms, which apparently had multiple sources
of invention, including psychometrics (Johnson
1967). They applied these simple tools to extract useful information
from, and to combine for the first time, enormous databases on molecular
pharmacology and gene expression (http://discover.nci.nih.gov/arraytools/).
They used two kinds of information from the drug discovery program of the
National Cancer Institute. The first kind of information described gene
expression in 1,375 genes of each of 60 human cancer cell lines. A target
matrix T had, as the numerical entry in row g and column c, the relative
abundance of the mRNA transcript of gene g in cell line c. The drug activity
matrix A summarized the pharmacology of 1,400 drugs acting on each of the
same 60 human cancer cell lines, including 118 drugs with “known mechanism of
action.” The number in row d and column c of the drug activity matrix A was
the activity of drug d in suppressing the growth of cell line c, or,
equivalently, the sensitivity of cell line c to drug d. The target matrix T
for gene expression contained 82,500 numbers, while the drug activity matrix
A had 84,000 numbers. These two matrices
have the same set of column headings but have different row labels. Given the
two matrices, precisely five sets of possible correlations could be
calculated, and Scherf et al. calculated all five. (1) The correlation
between two different columns of the activity matrix A led to a clustering of
cell lines according to their similarity of response to different drugs. (2)
The correlation between two different columns of the target matrix T led to a
clustering of the cell lines according to their similarity of gene
expression. This clustering differed very substantially from the clustering
of cell lines by drug sensitivity. (3) The correlation between different rows
of the activity matrix A led to a clustering of drugs according to their
activity patterns across all cell lines. (4) The correlation between
different rows of the target matrix T led to a clustering of genes according
to the pattern of mRNA expressed across the 60 cell lines. (5) Finally, the
correlation between a row of the activity matrix A and a row of the target
matrix T described the positive or negative co variation of drug activity
with gene expression. A positive correlation meant that the higher the level of
gene expression across the 60 cancer cell lines, the higher the effectiveness
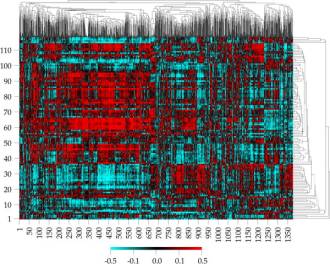
of the drug in suppressing the growth of those cell lines. The result of
analysing several hundred thousand experiments is summarized in a single
picture called a clustered image map (Figure 1). This clustered image map
plots gene expression–drug activity correlations as a function of clustered
genes (horizontal axis) and clustered drugs (showing only the 118 drugs with
“known function”) on the vertical axis (Weinstein
et al. 1997).
What use is this?
If a person's cancer cells have high expression for a particular gene, and
the correlation of that gene with drug activity is highly positive, then that
gene may serve as a marker for tumour cells likely to be inhibited
effectively by that drug. If the correlation with drug activity is negative,
then the marker gene may indicate when use of that drug is contraindicated. While important
scientific questions about this approach remain open, its usefulness in
generating hypotheses to be tested by further experiments is obvious. It is a
very insightful way of organizing and extracting meaning from many individual
observations. Without the microscope of mathematical methods and
computational power, the insight given by the clustered image map could not
be achieved. The Future To realize the
possibilities of effective synergy between biology and mathematics will
require both avoiding potential problems and seizing potential opportunities. Potential
problems. The productive
interaction of biology and mathematics will face problems that concern
education, intellectual property, and national security. Educating the next
generation of scientists will require early emphasis on quantitative skills
in primary and secondary schools and more opportunities for training in both
biology and mathematics at undergraduate, graduate, and postdoctoral levels (CUBE 2003). Intellectual
property rights may both stimulate and obstruct the potential synergy of
biology and mathematics. Science is a potlatch culture. The bigger one's gift
to the common pool of knowledge and techniques, the higher one's status, just
as in the potlatch culture of the Native Americans of the northwest coast of
North America. In the case of research in mathematics and biology,
intellectual property rights to algorithms and databases need to balance the
concerns of inventors, developers, and future researchers (Rai and Eisenberg 2003). A third area of
potential problems as well as opportunities is national security. Scientists
and national defenders can collaborate by supporting and doing open research
on the optimal design of monitoring networks and mitigation strategies for
all kinds of biological attacks (Wein et al.
2003). But openness of scientific methods or biological reagents in
microbiology may pose security risks in the hands of terrorists. Problems of
conserving privacy may arise when disparate databases are connected, such as
physician payment databases with disease diagnosis databases, or health
databases with law enforcement databases. Opportunities. Mathematical
models can circumvent ethical dilemmas. For example, in a study of the
household transmission of Chagas disease in northwest Argentina, Cohen and Gürtler (2001) wanted to know—since
dogs are a reservoir of infection—what would happen if dogs were removed from
bedroom areas, without spraying households with insecticides against the
insect that transmits infection. Because neither the householders nor the
state public health apparatus can afford to spray the households in some
areas, the realistic experiment would be to ask householders to remove the
dogs without spraying. But a researcher who goes to a household and observes
an insect infestation is morally obliged to spray and eliminate the
infestation. In a detailed mathematical model, it was easy to set a variable
representing the number of dogs in the bedroom areas to zero. All components
of the model were based on measurements made in real villages. The
calculation showed that banishing dogs from bedroom areas would substantially
reduce the intensity of infection in the absence of spraying, though spraying
would contribute to additional reductions in the intensity of infection. The
model was used to do an experiment conceptually that could not be done
ethically in a real village. The conceptual experiment suggested the value of
educating villagers about the important health benefits of removing dogs from
the bedroom areas. The future of a
scientific field is probably less predictable than the future in general.
Doubtless, though, there will be exciting opportunities for the collaboration
of mathematics and biology. Mathematics can help biologists grasp problems
that are otherwise too big (the biosphere) or too small (molecular
structure); too slow (macroevolution) or too fast (photosynthesis); too
remote in time (early extinctions) or too remote in space (life at extremes
on the earth and in space); too complex (the human brain) or too dangerous or
unethical (epidemiology of infectious agents). Box 1 summarizes five
biological and five mathematical challenges where interactions between
biology and mathematics may prove particularly fruitful. Acknowledgments This paper is
based on a talk given on February 12, 2003, as the keynote address at the
National Science Foundation (NSF)–National Institutes of Health (NIH) Joint
Symposium on Accelerating Mathematical–Biological Linkages, Bethesda,
Maryland; on June 12, 2003, as the first presentation in the 21st Century
Biology Lecture Series, National Science Foundation, Arlington, Virginia; and
on July 10, 2003, at a Congressional Lunch Briefing, co-sponsored by the
American Mathematical Society and Congressman Vernon J. Ehlers, Washington,
D.C. I thank Margaret Palmer, Sam Scheiner, Michael Steuerwalt, James
Cassatt, Mike Marron, John Whitmarsh, and directors of NSF and NIH for
organizing the NSF–NIH meeting, Mary Clutter and Joann P. Roskoski for
organizing my presentation at the NSF, Samuel M. Rankin III for organizing
the American Mathematical Society Congressional Lunch Briefing, and
Congressman Bob Filner for attending and participating. I am grateful for
constructive editing by Philip Bernstein, helpful suggestions on earlier
versions from Mary Clutter, Charles Delwiche, Bruce A. Fuchs, Yonatan Grad,
Alan Hastings, Kevin Lauderdale, Zaida Luthey-Schulten, Daniel C. Reuman,
Noah Rosenberg, Michael Pearson, and Samuel Scheiner, support from U.S. NSF
grant DEB 9981552, the help of Kathe Rogerson, and the hospitality of Mr. and
Mrs. William T. Golden during this work. Any opinions, findings, and
conclusions or recommendations expressed in this material are those of the
author and do not necessarily reflect the views of the NSF. References Anderson PW (1972) More is different. Science 177: 393–396. Find this article online Anderson PW (1995) Physics: The opening to complexity. Proc Natl
Acad Sci U S A 92: 6653–6654. Find this article online Bell ET (1937) Men of mathematics. New York: Simon and Schuster.
592 p. Benzer S (1959) On the topology of the genetic fine structure.
Proc Natl Acad Sci U S A 45: 1607–1620. Find this article online Bowers PL, Hurdal MK (2003) Planar conformal mappings of
piecewise flat surfaces. In Hege HC, Polthier K, editors. Visualization and
mathematics III Berlin: Springer. pp 3–34. Caswell H (2001) Matrix population models:
Construction, analysis and interpretation, 2nd ed. Sunderland
(Massachusetts): Sinauer Associates. 722 p. Cohen JE, Gürtler RE (2001) Modeling household transmission of
American trypanosomiasis [supplementary material]. Science 293: 694–698 Available: http://www.sciencemag.org/cgi/content/full/293/5530/694/DC1
via the Internet. Accessed 20 October 2004. Find this article online [CUBE] Committee on Undergraduate Biology
Education to Prepare Research Scientists for the 21st Century, Board on Life
Sciences, Division on Earth and Life Studies, National Research Council of
the National Academies (2003) BIO 2010: Transforming undergraduate education
for future research biologists. Washington (D.C.): National Academies Press.
191 p. Darwin F editor (1905) The life and letters
of Charles Darwin. New York: Appleton. Available: http://pages.britishlibrary.net/charles.darwin/texts/letters/letters1_02.html
via the Internet. Accessed 28 October 2003. Delwiche CF (1999) Tracing the web of plastid diversity through
the tapestry of life. Am Nat 154: S164–S177. Find this article online Delwiche CF (2000a) Gene transfer between organisms. In:
McGraw-Hill 2001 yearbook of science and technology New York: McGraw-Hill. pp
193–197. Delwiche CF (2000b) Griffins and chimeras: Evolution and
horizontal gene transfer. Bioscience 50: 85–87. Find this article online Delwiche CF, Palmer JD (1996) Rampant horizontal transfer and
duplication of rubisco genes in eubacteria and plastids. Mol Biol Evol 13:
873–882. Find this article online Erdös
P, Rényi A (1960) On the evolution of random graphs. Publ Math Inst Hung Acad
Sci 5: 17–61. Find this article online Euler L (1760) Recherches générales sur la mortalité
et la multiplication. Mémoires de l'Académie Royal des Sciences et Belles
Lettres 16: 144–164. Find this article online Ewens WJ (1972) The sampling theory of selectively neutral
alleles. Theor Popul Biol 3: 87–112. Find this article online Fisher RA (1937) The wave of advance of advantageous genes. Ann
Eugenics 7: 353–369. Find this article online Fisher RA (1950) Contributions to mathematical statistics. Tukey
J, indexer New York: Wiley. 1v. Galton F (1889) Natural inheritance. London: Macmillan. 259 p. Hamming RW (1971) Introduction to applied numerical analysis.
New York: McGraw-Hill. 331 p. Hardy GH (1908) Mendelian proportions in a mixed population.
Science 28: 49–50. Find this article online Hastings A, Palmer MA (2003) A bright future for biologists and
mathematicians? Science 299: 2003–2004. Find this article online Hastings A, Arzberger P, Bolker B, Ives
T, Johnson N, et al. (2003) Quantitative biology for the 21st century.
Available: http://www.sdsc.edu/QEIB/QEIB_final.html
via the Internet. Accessed 20 October 2004. Hsieh J (2003) Computed tomography: Principles, design,
artifacts, and recent advances. Bellingham (Washington): SPIE Optical
Engineering Press. 387 p. Hurdal MK, Bowers PL, Stephenson K, Sumners DWL, Rehm K, et al.
(1999) Quasi-conformally flat mapping the human
cerebellum. In: Taylor C, Colchester A, editors. Medical image computing and
computer-assisted intervention—MICCAI '99 Berlin: Springer. pp 279–286. Johnson SC (1967) Hierarchical clustering schemes. Psychometrika
2: 241–254. Find this article online Jungck JR (1997) Ten equations that changed biology: Mathematics
in problem-solving biology curricula. Bioscene 23: 11–36. Find this article online Kendall DG (1948) On the generalized birth-and-death process.
Ann Math Stat 19: 1–15. Find this article online Kendall DG (1949) Stochastic processes and population growth. J
R Stat Soc [Ser B] 11: 230–264. Find this article online Keyfitz N (1968) Introduction to the mathematics of population.
Reading (Massachusetts): Addison-Wesley. 450 p. Kimura M (1994) Population genetics,
molecular evolution, and the neutral theory: Selected papers. Takahata N,
editor Chicago: University of Chicago Press. 686 p. Kingman JFC (1982a) On the genealogy of large populations. J
Appl Prob 19A: 27–43. Find this article online Kingman JFC (1982b) The coalescent. Stoch Proc Appl 13: 235–248.
Find this article online Kolmogorov A, Petrovsky I, Piscounov N (1937) Etude de
l'équation de la diffusion avec croissance de la
quantité de matière et son application à un problème biologique. Moscow
University Bull Math 1: 1–25. Find this article online Levin S editor (1992) Mathematics and biology: The interface.
Challenges and opportunities. Lawrence Berkeley Laboratory Pub-701 Berkeley
(California): University of California. Available: http://www.bio.vu.nl/nvtb/Contents.html
via the Internet. Accessed 20 October 2004. Li L, Lindquist S (2000) Creating a protein-based element of
inheritance. Science 287: 661–664. Find this article online Liu JJ, Sondheimer N, Lindquist S (2002) Changes
in the middle region of Sup35 profoundly alter the nature of epigenetic
inheritance for the yeast prion [PSI+]. Proc Natl Acad Sci U S A 99:
16446–16453. Find this article online Lotka AJ (1925) Elements of physical biology. Baltimore:
Williams and Wilkins. 460 p. Luria SE, Delbrück M (1943) Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28: 491–511. Find this article online Margulis L, Sagan D (2002) Acquiring genomes: A theory of the
origins of species. New York: Basic Books. 240 p. Markov AA (1906) Extension of the law of large numbers to
dependent variables [Russian]. Izv Fiz-Matem Obsch Kazan Univ (2nd Ser) 15: 135–156. Find this article online Miller RG (1981) Simultaneous statistical
inference, 2nd ed. New York: Springer-Verlag. 299 p. Murray JA (1993) Mathematical biology, 2nd ed. Berlin:
Springer-Verlag. 767 p. Nicolson MH (1960) The breaking of the circle: Studies in the
effect of the “new science” upon seventeenth-century poetry,
revised ed. New York: Columbia University Press. 216 p. Orel V (1984) Mendel. Finn S, translator Oxford: Oxford
University Press. 111 p. Palmer MA, Arzberger P, Cohen JE, Hastings A, Holt RD, et al.
(2003) Accelerating mathematical-biological linkages.
Report of a joint National Science Foundation–National Institutes of Health
Workshop; 2003 February 12–13 National Institutes of Health. Bethesda,
Maryland Available: http://www.palmerlab.umd.edu/report.pdf.
Accessed 23 October 2004. Pearson K, Lee A (1903) On the laws of inheritance in man.
Biometrika 2: 357–462. Find this article online Provine WB (2001) The origins of theoretical population
genetics, 2nd ed. Chicago: University of Chicago Press. 211 p. Rai AK, Eisenberg RS (2003) Bayh-Dole reform and the progress of
biomedicine. Am Sci 91: 52–59. Find this article online Scherf U, Ross DT, Waltham M, Smith LH, Lee JK, et al. (2000) A
gene expression database for the molecular pharmacology of cancer. Nat Genet
24: 236–244. Find this article online Shipman JT, Wilson JD, Todd AW (2003) An introduction to
physical science, 10th ed. Boston: Houghton Mifflin. 1v. Smoller L (2001) Applications: Web-based precalculus. Did you
know…? Little Rock: University of Arkansas at Little Rock College of
Information Science and Systems Engineering. Available: http://www.ualr.edu/lasmoller/matrices.html
via the Internet. Accessed 26 December 2003. Turing AM (1952) The chemical basis of morphogenesis. Phil Trans
R Soc Lond B Biol Sci 237: 37–72. Find this article online Verhulst PF (1838) Notice sur la loi que la population suit dans
son accroissement. Correspondance mathématique et
physique publiée par A. Quételet (Brussels) 10: 113–121. Find this article online Volterra V (1931) Variations and fluctuations of the number of
individuals in animal species living together. In: Chapman RN, editor. Animal
ecology New York: McGraw Hill. pp 409–448. von Neumann J (1959) On the theory of
games of strategy. Bargmann S, translator. In: Tucker AW, Luce RD, editors.
Contributions to the theory of games, Volume 4 Princeton: Princeton
University Press. pp 13–42. von Neumann J, Morgenstern O (1953) Theory of games and economic behavior, 3rd ed. New York: John Wiley and
Sons. 641 p. Wein LM, Craft DL, Kaplan EH (2003) Emergency response to an
anthrax attack. Proc Natl Acad Sci U S A 100: 4346–4351. Find this article online Weinberg W (1908) Ueber den Nachweis der Vererbung beim
Menschen. Jahresh Verein f vaterl Naturk Württemb 64: 368–382. Find this article online Weinstein JN, Myers TG, O'Connor PM, Friend SH,
Fornace AJ, et al. (1997) An information-intensive approach to the molecular
pharmacology of cancer. Science 275: 343–349. Find this article online William H (1847) The works of William Harvey, M.D., physician to
the king, professor of anatomy and surgery to the College of Physicians.
Robert Willis. translator London: Printed for the
Sydenham Society 624 p. Yule GU (1925) The growth of population and the factors which
control it. J R Stat Soc 88: Part I 1–58. Find this article online
|